 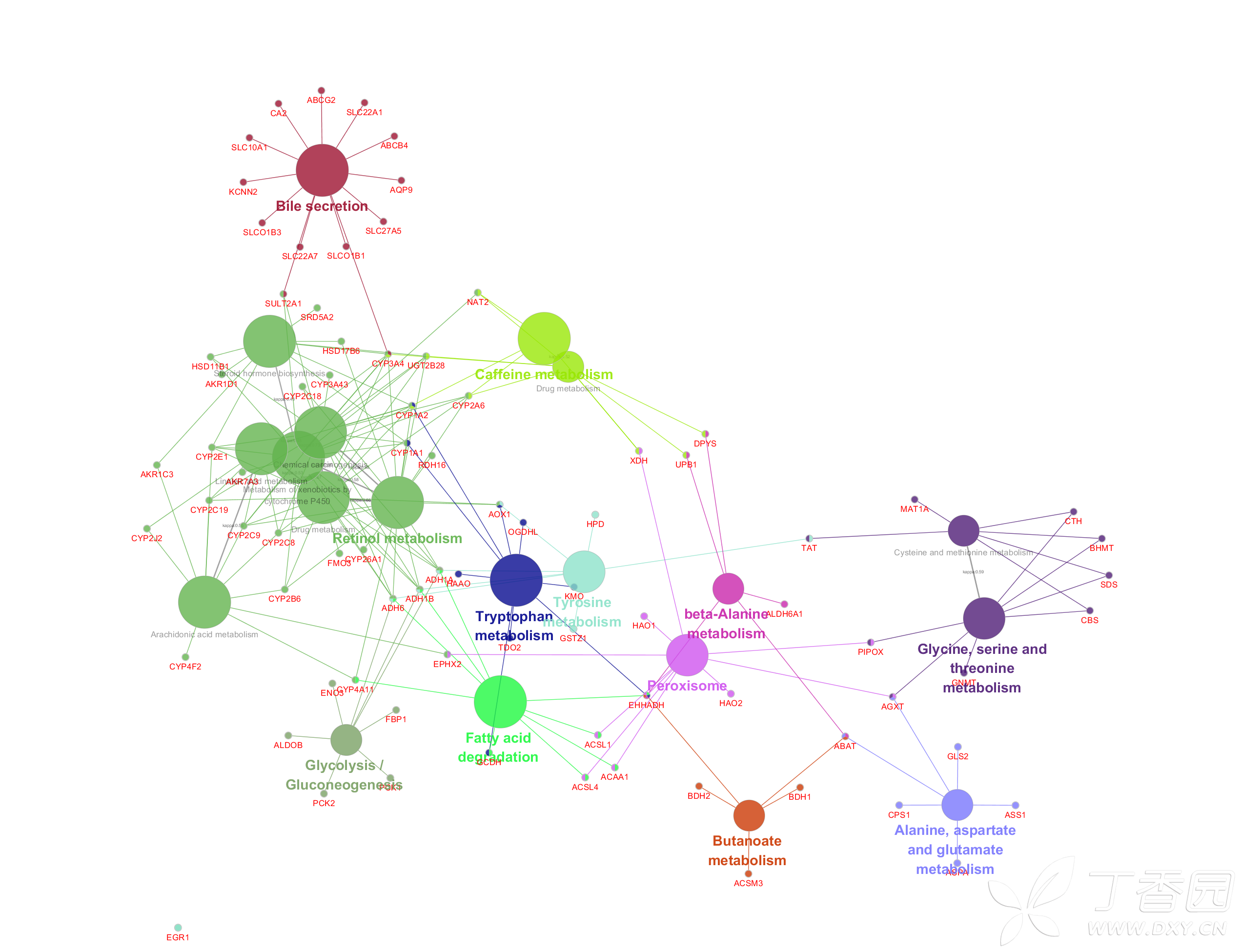
The newly found indirect interactions have been added in red. Left: Network representation of the identified pathīetween the two pathways, consisting of three proteins Gsk3b, Sgk3 and Tsc1. The comparison between HF and LF diet at t = 0. An indirect interaction between the Axon Guidance and Insulin Signaling pathways in the network for.Pathway interactions and what causes them.Non-redundant shortest paths in a weighted (2011) Exploring pathway interactions in insulin resistant mouse liver BMC Systems Biology 5: 127 Thomas Kelder, Lars Eijssen, Robert Kleemann, Marjan van Erk, Teake Kooistra, Chris Evelo Robert Caesar et al (2010) A combined transcriptomics and lipidomics analysis of subcutaneous,Įpididymal and mesenteric adipose tissue reveals marked functional differences. This allows for the detection of potential active regulatory mechanisms in the After loading interaction file(s), selecting a pathway element shows the interaction partners of thisĮlement and their expressions in a side panel. MiRNAs in pathways, which misses this regulatory information, and including all validated miRNA-target interactions, whichĬlutters the pathway. The Regulatory Interaction plugin offers a suitable middle-ground between not including any PathVisio RI plugin provides backpage info.Relationships, use BioPAX RDF 2 RDF 3 RDF 4 Kristina Hanspers, Bruce R Conklin, Chris T Evelo. Martijn P van Iersel, Alexander R Pico, Thomas Kelder, Jianjiong Gao, Isaac Ho, The BridgeDb Framework: Standardized Access to Gene, Protein and Metabolite Identifier Faculty of Health, Medicine and Life Sciences.BMC Bioinformatics 9: 399įaculty of Health, Medicine and Life Sciences Martijn P van Iersel, Thomas Kelder, Alexander R Pico, Kristina Hanspers, Susan Coort, Bruce R Conklin, ChrisĮvelo (2008) Presenting and exploring biological pathways with PathVisio. Can identify significantly changed processes.Uses gene expression, proteomics and metabolomics data.Data modeling and visualization on biological pathways.fumigatus’s metabolism to reveal dapsone, sulfamethazine, lovastatin and 3-bromopyruvic acid as potential drugs for the treatment of Aspergillosis. Our study provides a deeper insight into the mechanisms of A. Based on docking scores and MM-GBSA results, molecular simulations were carried out for 1AJ2-dapsone, 1DIS-sulfamethazine, 1T02-lovastatin and 70YL-3-bromopyruvic acid complexes, which validated our findings. Further, molecular docking and MM-GBSA analysis were performed with ligands chosen from DrugBank, and PubChem, and validated by experimental evidence and existing literature based on results from kinetic modeling and PPI network analysis. Based on the findings, dihydropteroate-synthase, dihydrofolate-reductase, 4-amino-4-deoxychorismate synthase, HMG-CoA-reductase, PG-isomerase and hexokinase could act as potential drug targets. For further analysis of the interaction of drug targets identified, a protein–protein interaction (PPI) network was built, and hub nodes were identified using the Cytohubba package from Cytoscape. While focusing on the folate biosynthesis, ergosterol biosynthesis and glycolytic pathway sensitivity, time-course and steady-state analysis were performed to find the proteins/enzymes that are essential in the pathway and can be considered as potential drug targets. 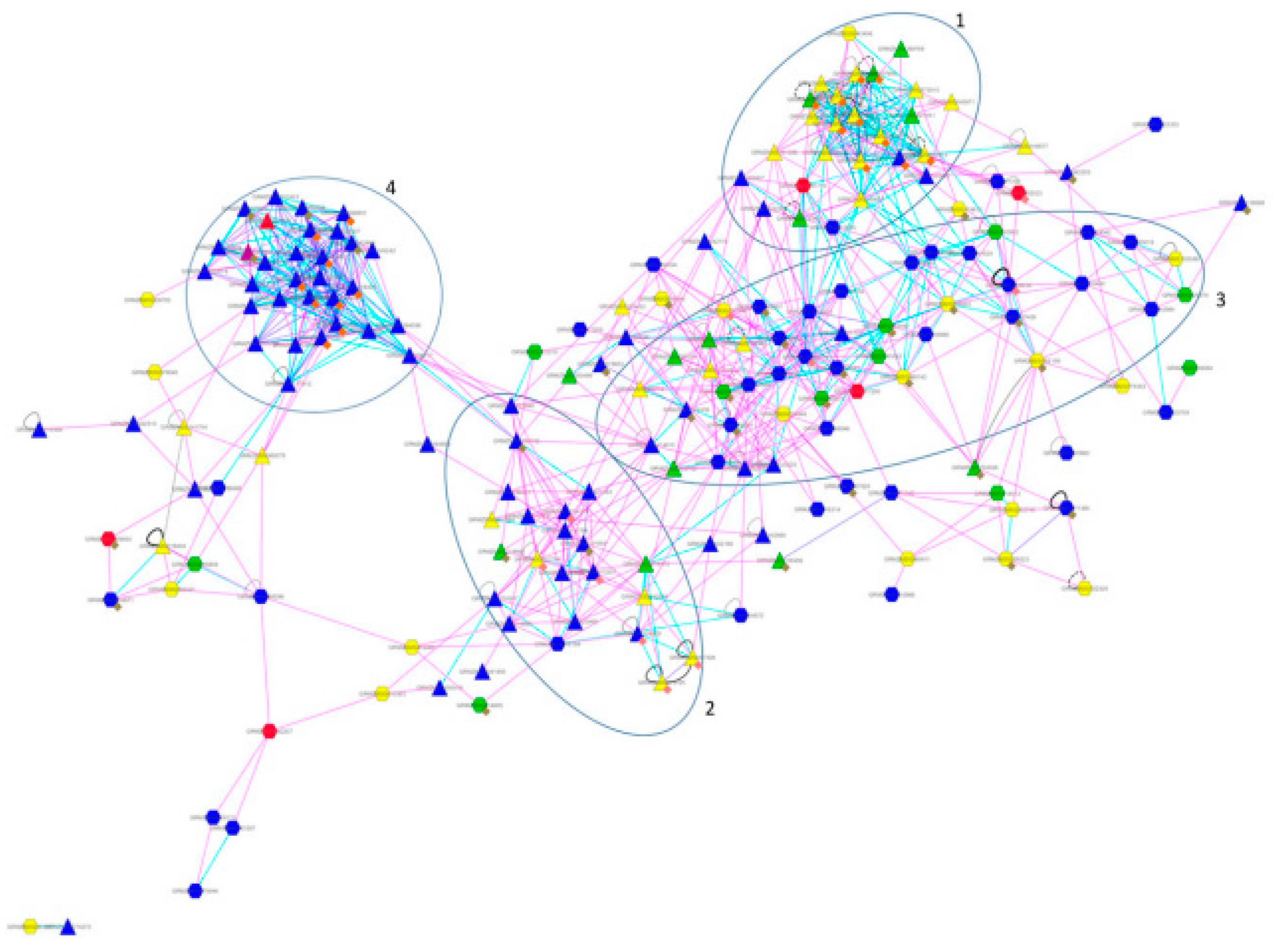
Our work focused on developing kinetic models of critical pathways crucial for the survival of A. To understand the pathogenicity of any organism, it is critical to identify the significant metabolic pathways that are involved. The diagnosis and treatment are difficult due to the diversity of individuals and risk factors and still pose a challenge for medical professionals. Aspergillosis is a major causative factor for morbidity in those with impaired immune systems, often caused by Aspergillus fumigatus. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
